Coronary Heart Disease (CHD), also called Atherosclerotic Heart Disease or Coronary Artery Disease (CAD) is a serious disease that affects millions of people around the world and that is caused by the formation of plaque in the arteries. There are two common approaches for plaque determination:
The present study focuses on the automatic segmentation (finding the Region of Interest or ROI) of stained microscopic images to analyze coronary plaque characteristics.
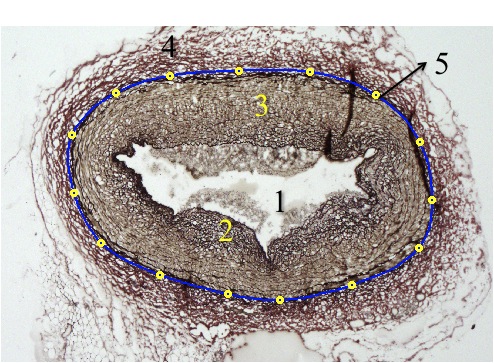
The artery
Basic parts of an artery, along with a brief description of each:
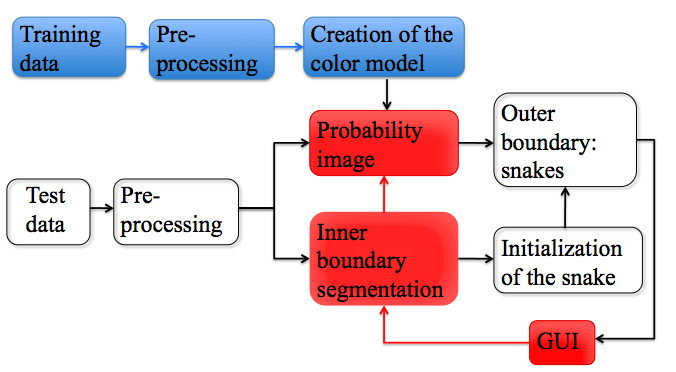
The basic structure of our algorithm is shown in Figure 1. In a timely fashion, we can divide the algorithm in:
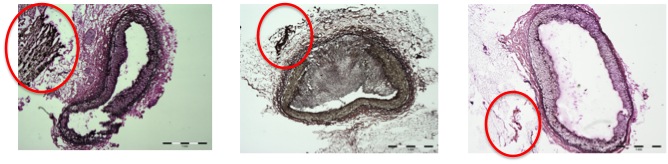 |
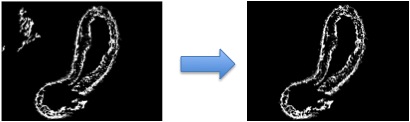

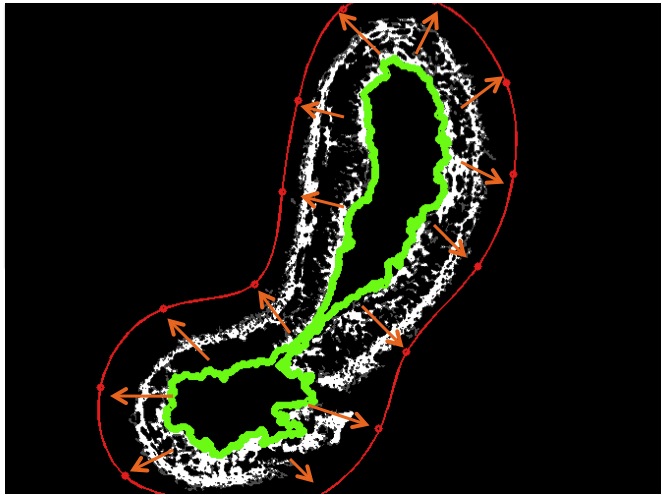 One of the most important steps to see our segmentation algorithm succeed, is finding a good initialization for the snake.
The initialization depends on the shape and size of the artery (problem 3), therefore we need to use known contour information. Our approach is to use the inner boundary found previously using region growing.
We start from the inner boundary (green) and dilate (orange arrows) the contour until we find that it is outside the EEL (red).
One of the most important steps to see our segmentation algorithm succeed, is finding a good initialization for the snake.
The initialization depends on the shape and size of the artery (problem 3), therefore we need to use known contour information. Our approach is to use the inner boundary found previously using region growing.
We start from the inner boundary (green) and dilate (orange arrows) the contour until we find that it is outside the EEL (red).
To analyze the performance of our automatic algorithm, we consider the manual segmentation of the images as ground truth.
We measure the error based on areas, comparing the number of pixels of the area found automatically and the number of pixels of the manually segmented area.
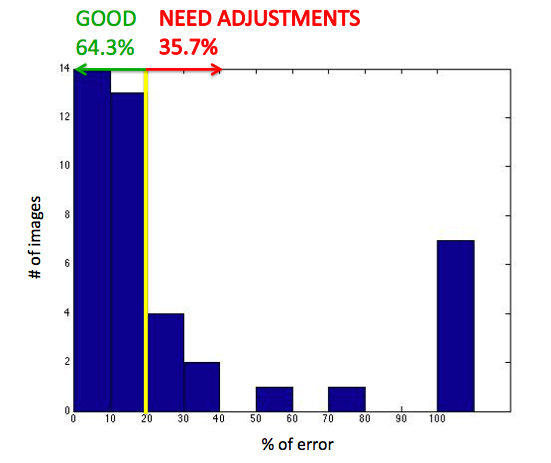
As we can see, we obtain statistically 54.8% of automatically segmented outer boundaries and 10% more with minimal user interaction.
From a user point of view, we can compute the average number of "mouse clicks" needed to segment an image as a measure of the workload of the user.
54.8% of the images are correctly analyzed with just one initial click (to choose the seed for region growing).
All the images can be analyzed with an average of 4.81 clicks.
SUMMARY
This project is focused on the problem of segmenting arteries in multi-stain histology images.
We propose an algorithm to automatically localize the artery and plaque within the image.
Our approach is based on the creation of a statistical color model from training data, which allows us to obtain likelihood images.
To find the outer boundary of the ROI in the likelihood images, we use snakes or active contours and minimize an error function which jointly optimizes a likelihood term, a smoothness term and a spline term. The initialization of the snakes is specially important to overcome the arteries shape differences and our solution is to use the inner boundary contour found by region growing.
The overall algorithm achieves positive results that specially reduce the amount of time that the user spends analyzing the images.
We obtain 64.3% of automatically segmented arteries.
If we consider an average of 20 clicks to segment each image manually and compare it with our algorithm that allows the user to segment all images with an average of 4.81 clicks, we see that we reduce the user's work by 76%.
In the future, we plan to exploit the difference in texture between the media and the adventitia to make our algorithm more robust.